Google Search: Keyword Search:
| Prev | ICM User's Guide 15.6 Atomic Property Field Tutorials | Next |
[ APF Superposition | Consensus Pharmacophore | APF Score | APF Screen | APF Cluster ]
APF Background
Atomic Property Fields (APF) is a 3D pharmacophoric potential implemented on a grid. APF can be generated from one or multiple ligands and seven properties are assigned from empiric physico-chemical components (hydrogen bond donors, acceptors, Sp2 hybridization, lipophilicity, size, electropositive/negative and charge).Here we describe template APF superposition whereby the APF is generated from a single or multiple template and is then globally optimized with the internal force-field energy of the ligand. You can read more about the APF method here.
About this Example In this example we will use Atomic Property Fields to superimpose, screen and cluster a set of Non-nucleoside reverse-transcriptase inhibitors (NNRTIs) (antiretroviral drugs) used in the treatment of human immunodeficiency virus (HIV). We will use a non-trivial superposition example of two NNRTIs which have similar pharmacophoric properties but are significantly different substructure. We will then read in additional PDBs which contain first generation NNRTIs approved by the FDA (Nevirapine, Delavirdine, Efavirenz) we will then use the combined fields to screen an sdf library to the APF fields
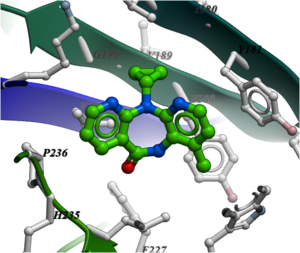
NNRTIs are chemically diverse but bind in the same allosteric pocket in the palm domain of the p66 subunit and part of the p51 subunit. Many NNRTIs bind in a "butterfly" conformation (see image below of Nevirapine) making pi-pi interaction with the rings in the pocket.
15.6.1 APF Superposition |
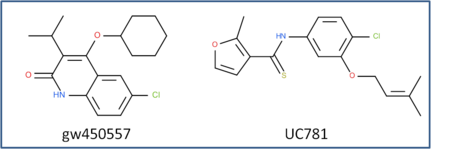
This is an example of a non-trivial chemical superpostion, we will superimpose inhibitor UC781 to gw450557. A prepared ICM file containing the two inhibitors can be downloaded here https://molsoft.com/addons/ftp/workshop (1. cut and paste the address into a web browser. 2. Right click on apf_superposition_example.icb and choose "Save Link As". 3. Open the .icb in ICM File/Open).
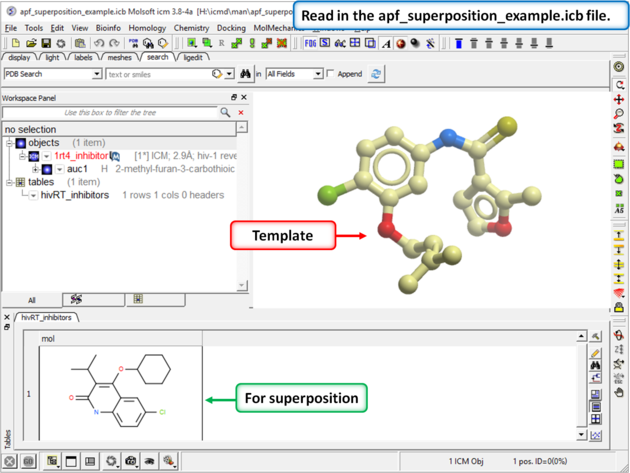 |
| Step 1: Download and open the prepared ICM file. The file can be downloaded here https://molsoft.com/addons/ftp/workshop (1. cut and paste the address into a web browser. 2. Right click on apf_superposition_example.icb and choose "Save Link As". 3. Open the .icb in ICM File/Open). |
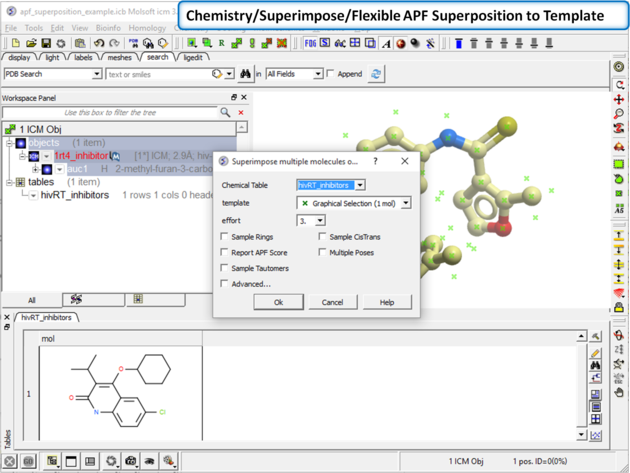 |
| Step 2: Superimpose the chemical from the table onto the 3D structure using Chemistry/Superimpose/Flexible APF Superposition to Template. Enter the parameters as shown in the screenshot. |
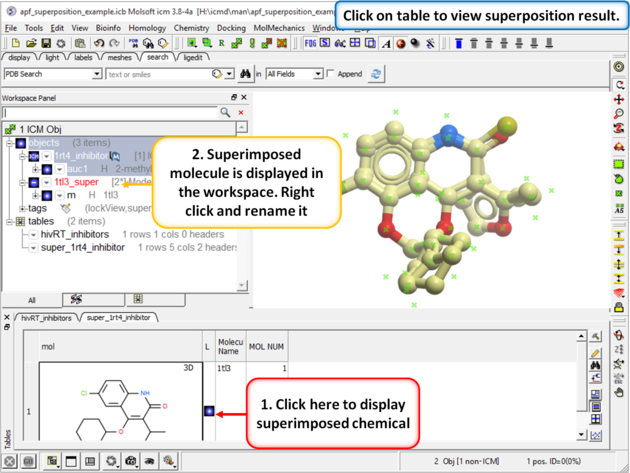 |
| Step 3: When the superposition has finished a results table will be displayed. You can toggle the superimposed molecule display on or off in the table. |
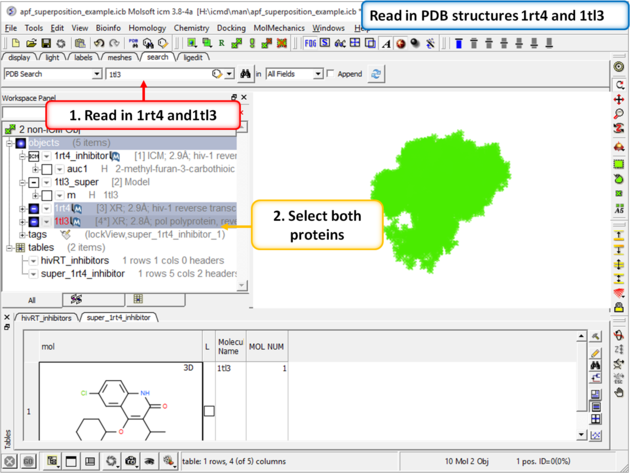 |
| Step 4: Now lets compare the docked pose of UC781 with the crystal structure pose. First we need to read in the PDB structures containing UC781 (1tl3) and gw450557 (1rt4). |
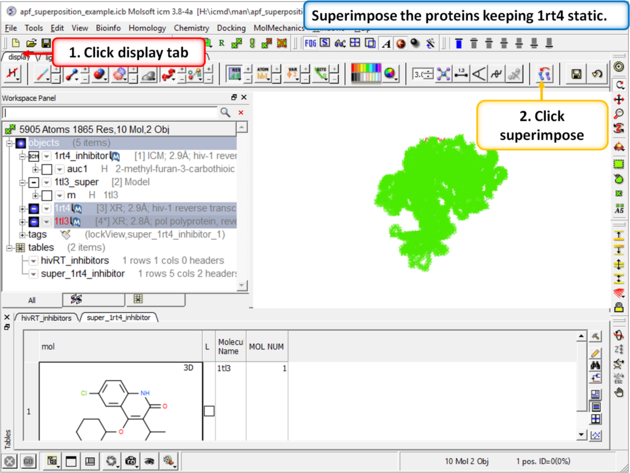 |
| Step 5: Superimpose the crystal structures keeping 1rt4 static. You need to select the two pdbs and use the superimpose button in the display tab. |
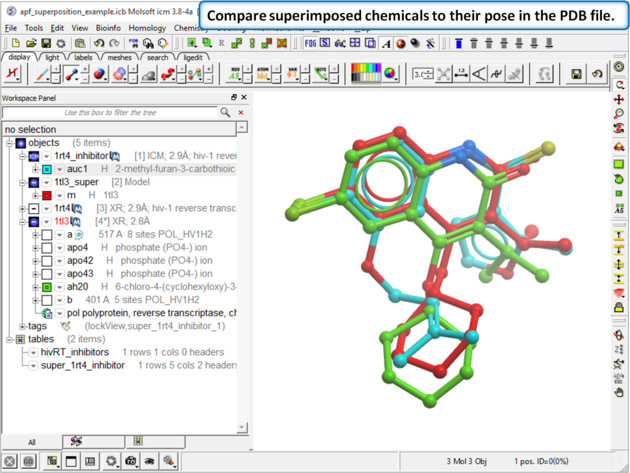 |
| Step 6: Compare the APF superimposed structure with the crystal structure. The RMSD should be <1.5. |
15.6.2 APF Consensus Pharmacophore |
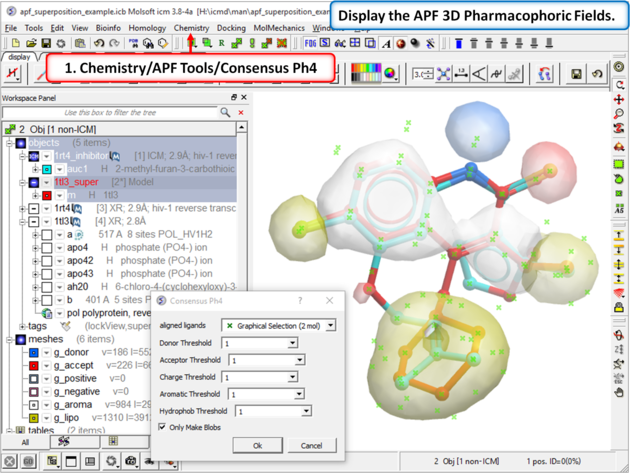 |
| Step 7: Display the APF consensus fields (Donor: blue Acceptor: red Aromatic: white Lipophyl: yellow Positive: lightblue Negative: pink). |
15.6.3 Calculate APF Score |
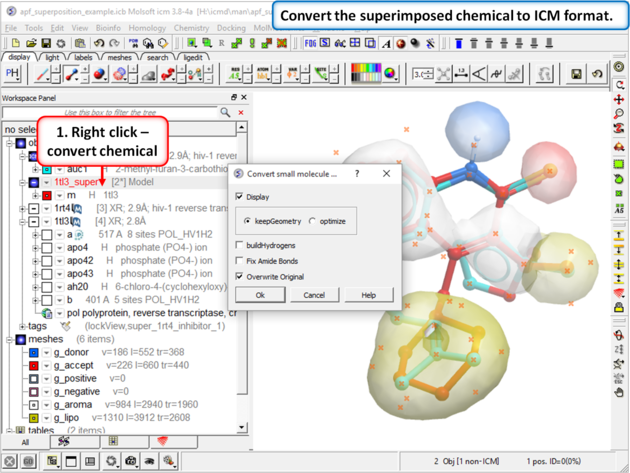 |
| Step 8: Convert the superimposed chemical to ICM format. |
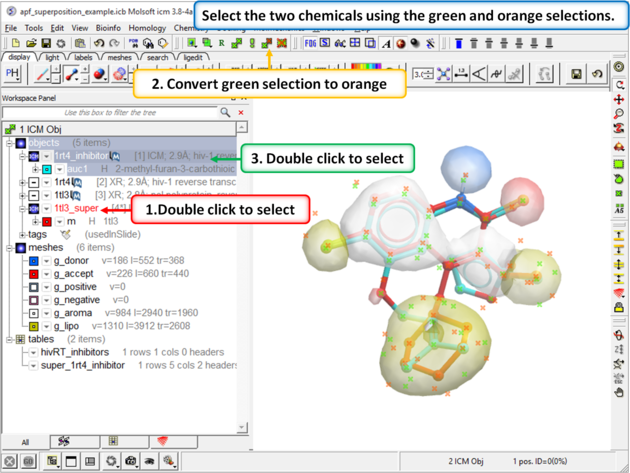 |
| Step 9: Select both the template and the docked ligand. |
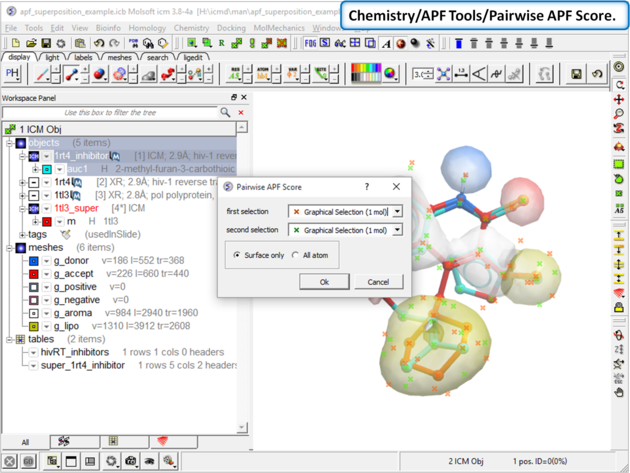 |
| Step 10: Choose the menu option Chemistry/APF Tools/Pairwise APF Score |
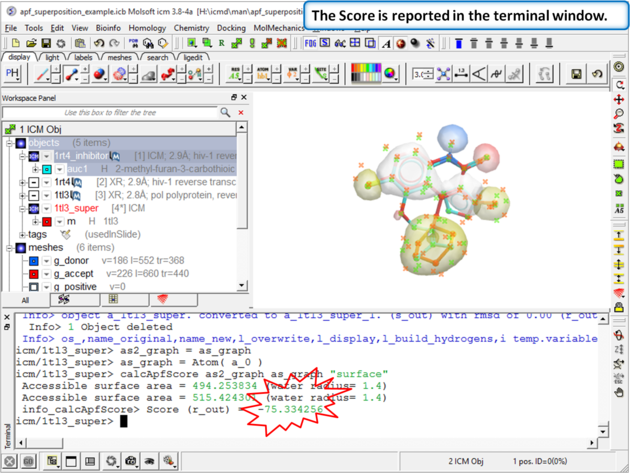 |
| Step 11: The score is reported in the terminal window (variable name r_out) |
15.6.4 Screen APF File |
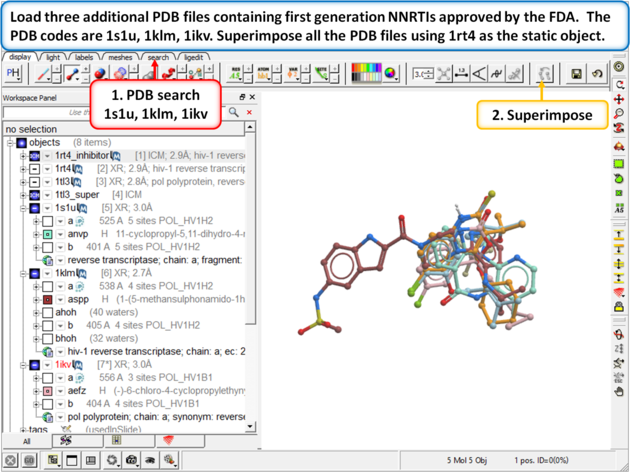 |
| Step 12: Load three additional PDB files containing first generation NNRTIs approved by the FDA. The PDB codes are 1s1u, 1klm, 1ikv. Superimpose all the PDB files using 1rt4 as static object. |
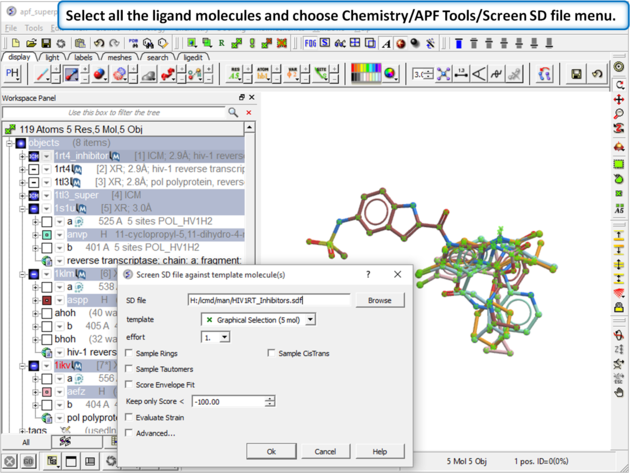 |
| Step 13: Select all the ligand molecules and choose Chemistry/APF Tools/Screen SD file menu. |
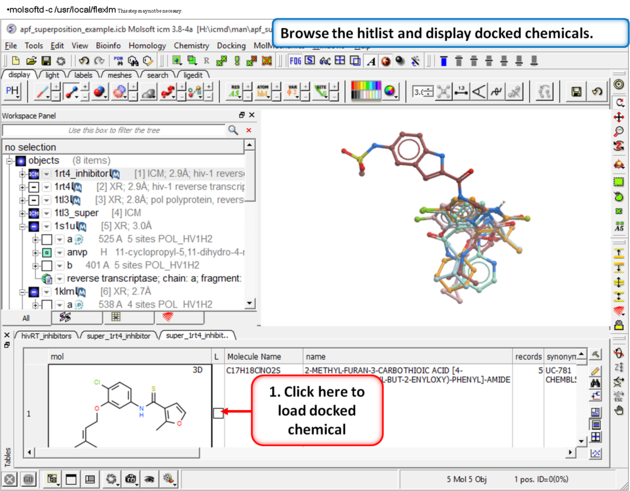 |
| Step 14: Browse the hitlist and display docked chemicals. |
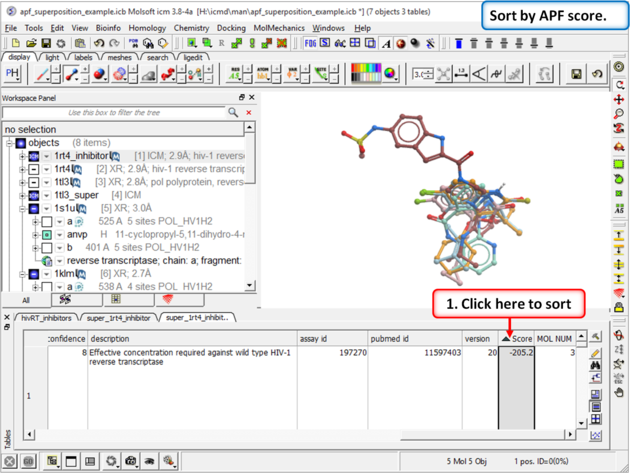 |
| Step 15: Sort the chemicals by SCORE and observe other NNRTI inhibitors in the hitlist |
15.6.5 APF Cluster |
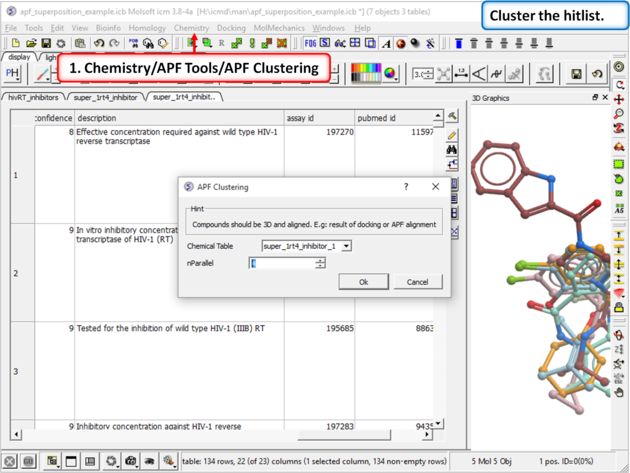 |
| Step 16: Cluster the hitlist. |
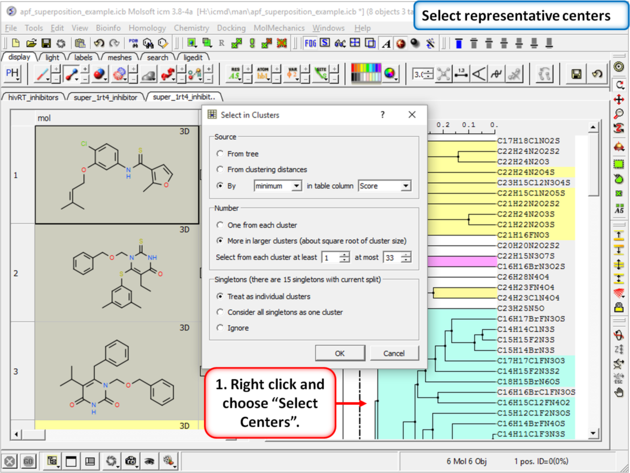 |
| Step 17: Select representative centers from the hitlist. |
| Prev 3D Ligand Editor Tutorial | Home Up | Next 2D QSAR Model |