| Prev | ICM User's Guide 8.1 Learn | Next |
[ About the Model | Model Weights | Improve Model | Save and Share Model ]
| 2D QSAR Tutorial |
To Learn from a data set and make a model:
First load in a table of data on which you wish to perform the learn and predict functions. See the tables chapter for more information on ICM tables.
- Read in your data e.g. chemical spreadsheet (e.g. File/Open SDF or .csv). It is usually recommended to Convert the IC50 to pIC50 (-LogIC50)
- Select Tools/Table/Build Prediction Model (Learn) and a window or use the Chemistry/Build Prediction Model option.
- Enter the name of table with which you want to perform the predictions. You may locate your table from the drop down arrow menu.
- Select the Column from which you wish to learn. Use the drop down arrow to select. NOTE If the table does not contain any numeric (integer or real) columns, there is nothing to predict, so the "Build Prediction Model - Learn" button will be disabled.
- Enter a name for the learn model.
- Select which learning method you wish to use from the drop down menu (PLS, PCRegression, Bayesian Classifier, Random Forest).
- Select whether you would like ICM to estimate the number of latent variables or you can specify a range.
- Select No Free Term Constraint:f(0)=0 if you would the PLS method to find a constant and add it to the model.
Descriptors
- Select which columns (descriptors) of your table you wish to use to 'learn'. You can select just the mol column or all numerical columns.
- If you are using chemical desriptors select the fingerprint method want to use Linear or Extended Connectivity Fingerprints (ECFP)
- Select Binary if you want to use Binary fingerprints - do not check if you want to use counted.
- Minimal chain length the minimal length of the chain of atoms, enumerated in each compound, usually it is 1 which means that you will consider an atomic composition numbers as descriptors. The typing of atoms can be customized with the map argument. .
- Maximal chain length the maximal length of the chain atoms and bonds enumerated in each compound Usually it is 3 or 4. Larger values of this parameter will lead to a large number of possible combinations. To overcome that, either the typing needs to be simple (e.g. only sp2 and sp3 property, regardless of the atom number), or the data set needs to be really large.
- Length - this is the total number of bins in a final fingerprint. Typically the size (either explicit or automatically estimated) is in a hundreds to throusands range. We recommend to check the auto option.
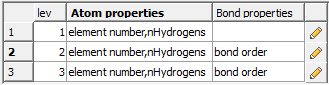
- In this dialog you customize the atom properties for every chain length or ecfp fragment size. Click on the "pencil button" to add additional properties to a chain or fragment size (lev). Once the model is made you will see the weights of these properties in the model (right click on model in the ICM workspace and choose Weights).
Test
- Select the number of cross-validation groups you wish to use or selected rows can be used for cross validation. The number of iterations will impact the speed of the calculation. 5 is the default number of groups but 2 would be the least rigorous and selecting the 'Leave-1-out' would be the most rigorous calculation.
- Bootstrapping generates statistics (R2, RMSD) for a random prediction and you should aim for there to be a significant difference between the random R2 and the model R2.
- Click on the learn button and a model will be created and placed in the ICM Workspace panel (left hand side).
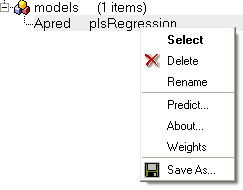
All models are then stored in the ICM workspace as shown below. A number of options are displayed in the right click menu.
8.1.1 About the Model |
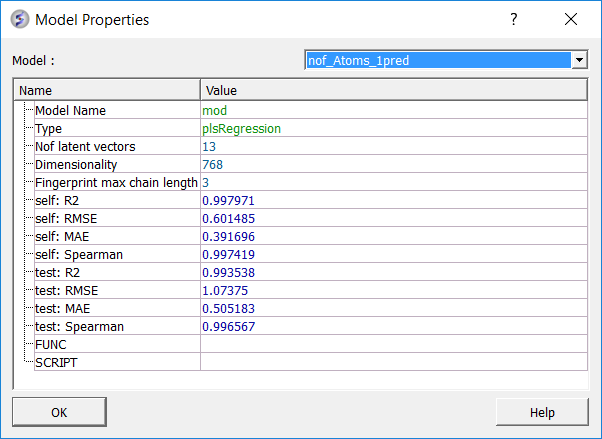
To see more details about your model (e.g. R2, RMSE etc....):
- Right click on your model in the ICM workspace and choose Model Data.
8.1.2 Model Weights |
The Weights dialog can be helpful to see what is driving the correlation in your model or for troubleshooting. The key column is "w" which represents the weight of the fragment in the regression. You may see a particular fragment providing a high weight to the model so you may want to add similar fragments to your training set to see if it will improve the model. Removing fragments from the training set based on their weight is not recommended because each fragment is part of a multiple component and it is difficult to know precisely its importance.
- Right click on the model in the ICM Workspace and choose Weights
- A table will be displayed containing the following columns:
name = smiles string of fragment. You can copy and paste this into the molecular editor to view it in 2D.
w = is actual weight of the fragment in the regression (used directly to get the result value).
mean = indicates the mean occurence of the fragment in the model . For example, a value of 6 means the fragment is used 6 times in the regression.
rmsd = a high value indicates high occurence of the fragment and influences the relative weight parameter.
wRel = relative 'importance' of the fragment based on the RMSD, mean and CorrY values.
8.1.3 Improve Model |
The model can be improved by:
- Right click on the model in the ICM Workspace and choose Improve
- A dialog box similar to the one you used to make the model in Learn will be displayed.
8.1.4 Save and Share Model |
You can save and share a model by:
- Right click on the model in the ICM workspace (left hand panel) and choose Save As... This will save the model in MolSoft's .icb format.
- The .icb file can be opened in another session using File/Open
| Prev Learn and Predict | Home Up | Next Predict |