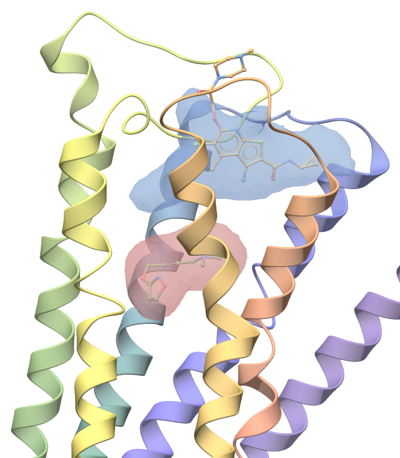
ICM Pocket Finder is a method which will identify ligand binding sites in any protein, DNA or RNA structure. Protein-ligand binding sites are identified based on the grid potential map of van der Waals interaction of the receptor and the surfaces are contoured. Physical properties of the pockets are then calculated and tabulated and a drug-likeness score is provided.
About the Method The ICM Pocket Finder method (1-2) uses only the protein structure for the prediction of cavities and clefts. No prior knowledge of the substrate is required. The position and size of the ligand-binding pocket are determined based on a transformation of the Lennard-Jones potential by convolution with a Gaussian kernel of a certain size, a grid map of a binding potential and construction of equipotential surfaces along the maps. The pockets are displayed graphically as a surface and the dimensions of each pocket are presented in an interactive table and plot.
Factors that can influence ligand binding to a pocket include the pocket volume and area, buriedness, hydrophobicity, and how compact the pocket is. All these properties are calculated using ICMPocketFinder and tabulated. Scientists at Merck used MolSoft.s ICMPocketFinder algorithm to define a way for quantifying "drugability" of a protein target (3). The metric they used is called Drug-Like-Density (DLID), and this score is also provided in the results table. A good example of the use of the icmPocketFinder method is in the database called Pocketome. The Pocketome (www.pocketome.org) is an encyclopedia of conformational ensembles of all druggable binding sites that can be identified experimentally from co-crystal structures in the Protein Data Bank (4).