[ Load Models | Index Models | Run MolScreen ]
For a full panel of models a MolScreen license is needed. A small subset of models are provided free of charge Click here to see the available models and model types.
To load the models:
- Select MolScreen Model/Load Indivifual Pre-defined Model Panel.
- Select the directory containing your models. By default they will be in the .icm/models directory but you can find the directory location by going to File/Preferences and checking the directories tab and look for Tools.models Directory.
- Select "All Targets" to load all models or use the drop down button to select a subset of models.
- This will load a table listing the available models - each model is prefixed by the model type. Each model is in a row in the table - to view the performance of a model select the model by selecting the row number and then choose Check Model Performance in the Tools panel of the table.
This option is only needed if you have built custom models that you want to include into MolScreen of if the ICM version or model directory has been updated.
- Select MolScreen Model/Index All Model Types.
To Run MolScreen
- Read in a chemical table (sdf file) you want to screen.
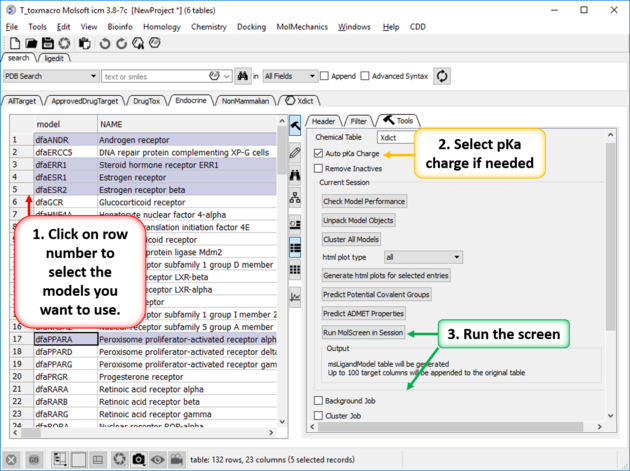
- Select one or more models in the model panel table. You can select them by clicking on the row number - the row should then be highlighted. Range of models can be selected by clicking and holding the shift key - non-contiguous selections can be made by clicking and holding the CTRL key.
- If you want to use MolSoft's built in pKa model check the box labeled Auto pKa Charge.
- If you want to display only the results with active hits check the box labeled Remove Inactives.
- If you have a small set of chemicals and number of models you can run the screen by clicking on Run MolScreen in Session. If you have larger sets you should choose the option Background Job and enter the number of processors you want to use. The jobs can also be sent to a cluster - to do this check option Cluster Job and enter how you want to split the job and the server details.