Google Search: Keyword Search:
| Prev | ICM User's Guide 5.3 Protein Superposition | Next |
[ Display and Select Proteins for Superposition | Superimpose Button ]
| Available in the following product(s): ICM-Browser-Pro | ICM-Pro |
 One or more proteins can be superimposed. Simply select the molecules or parts of the molecules you wish to superimpose and then use the selection of
protein superimpose tools described in this section. A convenient superimpose button can be found in the Display tab (see image of button (left).
One or more proteins can be superimposed. Simply select the molecules or parts of the molecules you wish to superimpose and then use the selection of
protein superimpose tools described in this section. A convenient superimpose button can be found in the Display tab (see image of button (left).
Chapter Contents:
- Select Proteins for Superposition
- Superimpose Button
- Superimpose by 3D
- Superimpose Multiple Proteins
- Arrange as Grid
- Superimpose Sites by Atomic Property Fields
5.3.1 Select Proteins for Superposition |
Before any superposition operation can be undertaken you need to select the protein structures you wish to superimpose.
One way to do this is by selecting in the ICM workspace. For other selection tools please see the Making Selections section of the manual.
- Select both receptors by double clicking on the name of the molecule in the ICM Workspace. To select two molecules use the Ctrl button or use the shift button to select a range of objects in the ICM Workspace. A receptor which is selected will be highlighted in blue in the ICM Workspace and with green crosses in the graphical display.
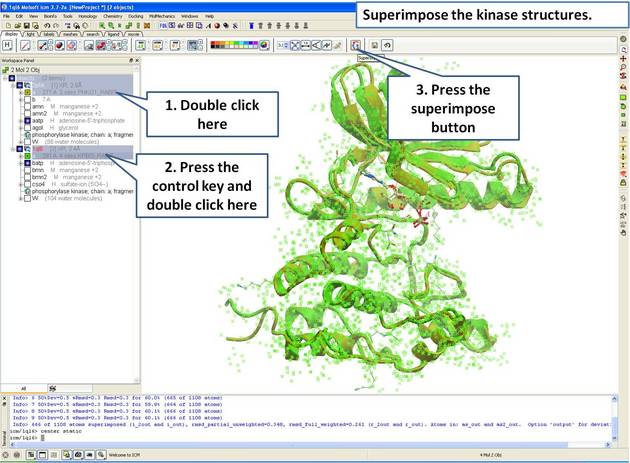
Once the molecules are selected you can then superimpose them using the options described in the next section of this manual.
5.3.2 Superimpose Button |
A convenient way to superimpose two molecules is by using the superimpose button in the display tab, ICM will calculate the Ca-atom, backbone atom and heavy atom differences between the two structures. More advanced superimpose options can be found in the Tools/Superimpose menu.
To superimpose:
- First load the two structures into ICM.
- Select which parts or all of the two structure you wish to superimpose (see the chapter on Selections or the description protein-superposition-select{here}.).
- Select the display tab (previously called Advanced tab) at the top of the GUI.
- Select the superimpose button.

The rmsd will be displayed in the terminal window as shown below:
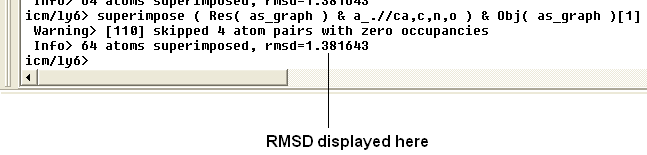

| NOTE: You do not need to select the whole molecule, the superimpose button will work on small selections e.g the loop regions or domains. |
| Prev PiPi | Home Up | Next 3D Similarity |