[ Chemical Search | Molecular Editor | convert to 3D | Grid View | Convert Chemical | convert to 3D and preserve | Add Property Column | Extract Ligand | Druglikeness | Chemical Monitor | SMILES | APF Model | Chemical Rotate | Remove salt | Fingerprints | Distance to Ring | H-bond acceptors ]
Frequently asked questions regarding small molecules, ICM-Chemistry tools and MolCart
- How do I perform a chemical search?
- How can I draw small molecules?
- How do I read in a small molecule from ISIS draw and convert it to 3D?
- How can I change the layout of a chemical table?
- How can I convert a chemical in a chemical table into 3D?
- I have a small molecule which already has the 3D coordinates defined. How can I load the molecule and not optimize it so as to preserve the assigned 3D coordinates?
- I have a chemical table displayed - how can I add columns of chemical properties associated with each chemical in my table?
- I have a small molecule displayed in 3D in a loaded PDB file. How can I extract this molecule into an ICM Chemical Table?
- What is considered a good druglikeness value?
- I do not see the chemical property monitor in the molecular editor. Where is it?
- How do I convert SMILES string into a 2D structure
- Is there a way to build a classification model using the APF output?
- How to rotate a 2D chemical sketch so it fits nicely in its cell in a chemical table?
- How do I remove unwanted metal and counterions from an SDF file?
- How to export binary fingerprints?
- How to measure the distance to a center of a ring?
- How does ICM calculate the number of hydrogen bond acceptors in a chemical?
You can search MolCart or Chemical Tables using the ICM Chemical Search Window. This window can be displayed by going to:
- Tools/Chemistry/Chemical Search
OR
- Click on this icon
OR
- Right click on a structure in a chemical table and select Query Molecule.
OR
- Or right click on a database in MolCart you wish to search and select Query...
Use the molecule editor
Chemistry/Molecular Editor
or look for the ICM molecular editor button at the top of the graphical user interface.
- Save the molecule in mol format and then read into ICM (File/Open).
- The molecule should be displayed in 2D in a molecular table.
- Right click on the molecule in the table and select chemistry/convert to 3D and optimize.
| NOTE: There is no need to use an external chemistry drawing software when you can use the ICM molecular editor which is fully integrated into the ICM software. |
To change the layout of a chemical table (eg converting a table to grid view).
- Select the columns you wish to display in grid view. You can do this by clicking on the column headers with the CTRL key pressed down.
- Once the columns are selected right click inside the table and select Table View
- Select Custom Grid...
To do this:
- Select the chemical or chemicals you wish to convert to 3D. You can do this by clicking on the row number whilst keeping the CTRL key pressed down.
- Right click on the chemical table and select Chemistry/Convert 3D and Optimize
- File/Open and read in the molecule - it should be then displayed in a molecular table
- Right click on the molecule in the table and select Chemistry/ Load and Preserve Coordinates.
To read a chemical table into ICM:
- File/Open and look for sdf files.
To add a chemical property to the table.
- Right click on the 'mol' column header and select Insert Column...
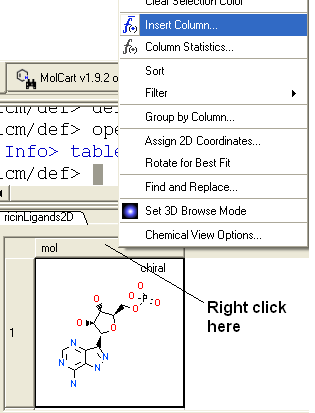
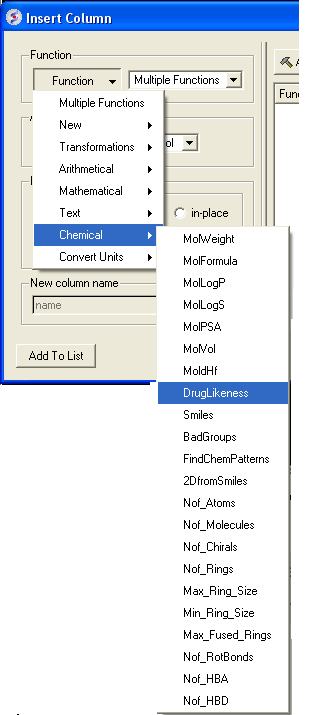
- Click on the drop down 'Function' button and select chemical.
- Select which property you wouls like to add and click OK.
- The property you selected will be displayed in the table.
You can extract a ligand from an ICM object or PDB file by:
- Right click on the ligand in the ICM Workspace.=
- Select Extract Ligand.
- Choose to extract either 2D or 3D coordinates and the molecule will be placed in a chemical table.
When building a molecule in the ICM Molecular Editor (Tools/Chemistry) properties such as druglikeness are calculated on the fly. The properties can also be added by inserting a column into a chemical table (right click on column header/ insert column/ Function = Chemical). These values should be used as a guide and druglikeness is a prediction based on drug-like properties. A druglikeness value less than zero indicates that the compound may have some non-drug-like properties.
If you do not see the chemical monitor in the ICM Molecule Editor - Go to:
- Go to Molecular Editor
- View/Chemical Monitor
See Convert Smiles to 2D section.
_setAPFparams is in the distribution since 3.6-0;
The usage:
icm _apf3Dqsar train=trainingSet.sdf activity=LogIC50 table=testSet.sdf
Training and test set compounds should be all pre-aligned, for example by aligning training set actives using APF multiple chemical alignment, and then superimposing the test compounds onto aligned actives using APF superposition. Any external alignment method can be used as well. The field containing activity data in the training set SDF is specified by activity= argument.
The script also can take alignments in icm multiple object format *.ob, in which case SDFs are only used for input/output of activity data and can be just 2D:
icm _apf3Dqsar train=trainingSet.sdf align=trainingSet3Daligned.ob activity=LogIC50 predict=testSet3Daligned.ob table=testSet.sdf
The results are written to testSet_predict.sdf output file. Some statistics is reported along the way. If testSet.sdf contains activity (i.e. LogIC50) column like the training set, RMSD and R2 will be reported as well.
See this description in the command line manual:
http://www.molsoft.com/man/icm-commands.html#make-flat
You can do this in the menu option Chemistry/Standardize and check the remove salt option.
To export our default binary fingerprints the Descriptor function can be used as below:
Sarray(Descriptor(t.mol))
It will return array of binary 0/1 strings of length 1536.
If you need to just measure the distance from the center of the ring to the atom it can be done lile this:
ring atoms are selected into as_graph and the distance is measured from the center of the mass to atom 'c3'
Distance( Matrix( Mean(Xyz(as_graph ) ) ) Xyz( a_//c3 ) )
to visualize this you can create a dummy atom in the center of the ring
# create a single atom with coordinates in the center of the ring
read mol Chemical( "C|3D:" + Sum(Sarray( Mean(Xyz(as_graph )))",") ) name="cent_ring"
# display it
display xstick a_cent_ring.m
# display distance
make distance append refresh a_cent_ring.m/1/c1 a_//c3 display
The number of hydrogen bond acceptors is calculated as a sum of number of lone pairs on the following atoms:
- any oxygen (double bonded - 2 pair, single bonded - 1 pair)
- SP1 nitrogen (1 coordinated) - 1 pair
- SP2 nitrogen (2 coordinated) - 1 pair
- SP3 nitrogen (3 coordinated) - 1 pair
- 2 valency sulfur (same as oxygen)