| Available in the following product(s): ICM-Chemist | ICM-Chemist-Pro | ICM-VLS |
Chemicals displayed in an ICM Molecular Table can be split into fragments. This is useful for generating a series of R-groups to be added to a scaffold (See section describing reactions.
To generate fragments:
- Select the column or row(s) you wish to generate the fragment from.
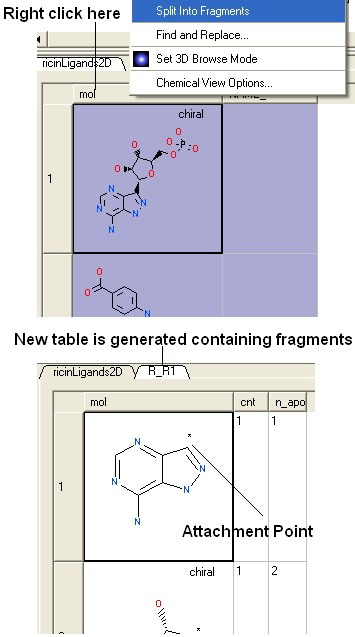
- Right click on the "mol" column header and select "Split Into Fragments".
- A new table of chemical fragments will be displayed. Each fragment is assigned an attachment point which is flagged with an asterisk (*).