| Available in the following product(s): ICM-VLS | |
This option allows you to perform a ligand-based screen using the Atomic Property Fields (APF) of superimposed chemicals. SD files are screened and the ligands are scored according to their fit into the APF field. Read more about APF here: http://www.molsoft.com/apf.html
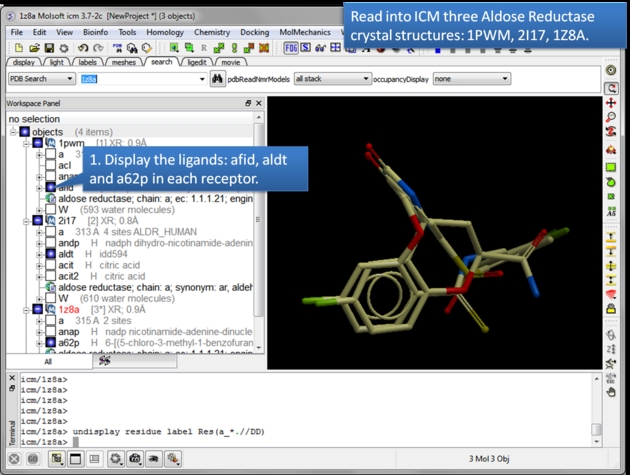 |
| Read in and display the superimposed template chemicals. In this example we will use three aldose reductase crystal structures 1PWM, 2I17, and 1Z8A. IMPORTANT: Convert each PDB file to an ICM object. |
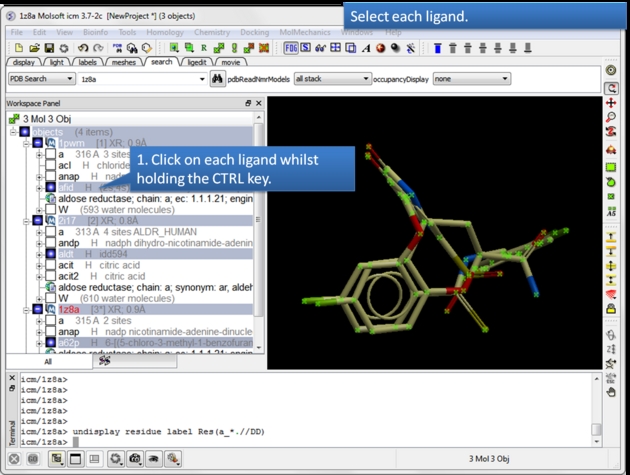 |
| Select each chemical you wish to contribute to the APF fields. Select by double clicking on the chemical in the ICM workspace and hold down the CTRL key. |
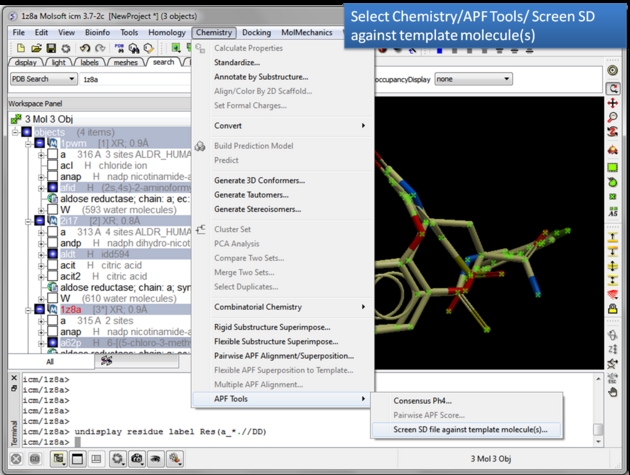 |
| Choose the Screen SD file against template molecule(s) option. Chemistry/APF tools/Screen SD file against template molecule(s) |
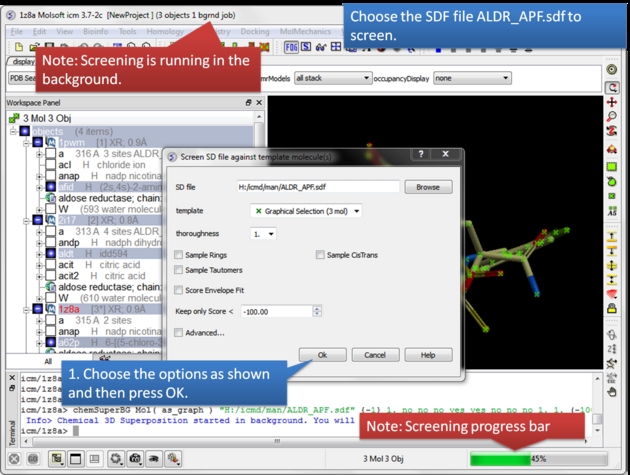 |
Select the SD database you wish to screen.
|
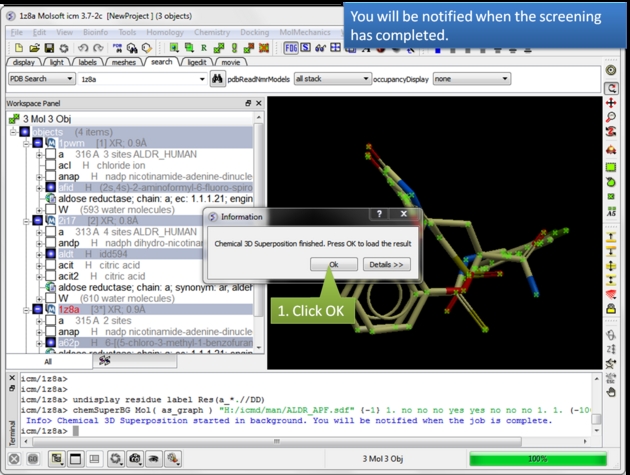 |
| A notification will be displayed when the screen is completed.Click OK and a hitlist table will be displayed. |
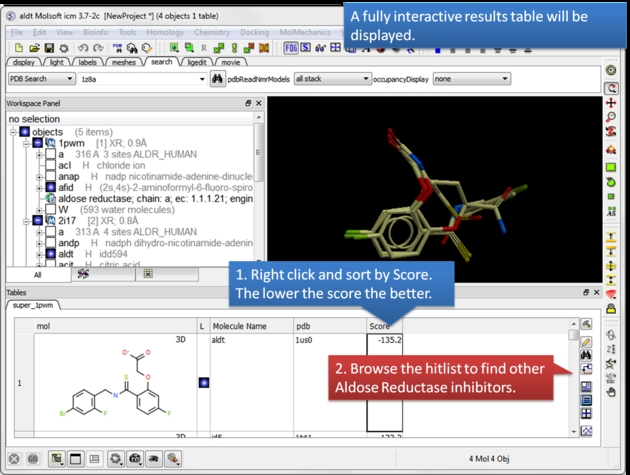 |
| A hitlist table will be displayed. You can sort the table by score and toggle the display on and off using the buttons in the L column. |
Guide to the RIDE Results:
- Score: The Score is the sum of all atomic contributions to the overall Atomic Property Field (APF) score, plus any penalties (e.g., envelope or excluded volume). Each atom is assessed by how well its properties (such as hydrogen bonding, charge, hydrophobicity, aromaticity, and size) match the target environment. A lower score indicates a better fit.
- Strain: strain in kcal/mol - the lower the better
- NormScore: normalized by self Score. values: [0-1]