Google Search the Manual:
Keyword Search:
| Prev | ICM User's Guide 22.2 Drug Target Analysis and Preparation | Next |
Drug Target Analysis & Preparation in ICM-Pro
Watch the Full Video: Drug Target Analysis and Preparation in ICM-Pro
PDB to ICM Conversion
Objective: convert the PDB to a full atom model for modeling and docking.
- Read in PDB > Search Tab > Enter "1xbb"
- Conversion Protocol: Right-click the object >
Convert PDB[00:01:43] - Atom Completion: ICM automatically builds missing heavy atoms in side chains and adds hydrogens based on internal residue libraries [00:00:22]
- Force Field Assignment: Assigns MMFF atom types, partial charges, and formal charges essential for calculating binding energy [00:00:52]
- Optimization:
- Hydrogen Network: Global optimization of H-bonding networks [00:00:28]
- Residue Flips: Asparagine (Asn) and Glutamine (Gln) are flipped to maximize H-bond potential; Histidine (His) protonation is determined by the local environment [00:00:38]
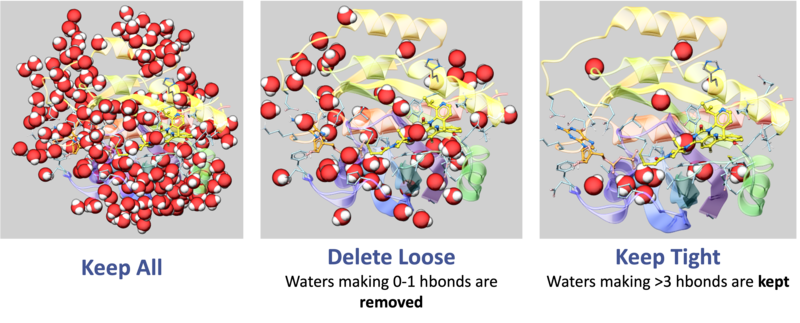
- Water Management: Retain "Tight" waters (>3 H-bonds) to maintain structural integrity of the pocket; delete "Loose" waters (0-1 H-bonds) to prevent artificial steric hindrance during docking [00:01:28]

II. Structural Quality Control (QC)
Objective: Quantify the reliability of the atomic coordinates.
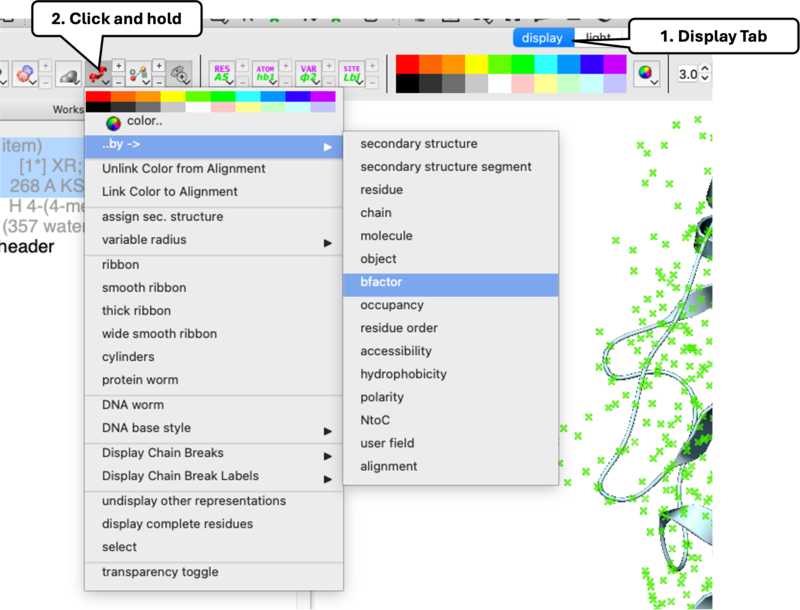
- B-Factor: Navigate to
Display> click-holdRibbon>Color by B-Factor[00:03:26]. Warmer colors indicate higher B-Factor greater flexibility. - Occupancy: Double-click ligand to select it > click-hold
Stickin Display Tab >Color by Occupancy[00:03:57]. Low occupancy indicates unreliable atom placement.
III. Electron Density Map Validation
Objective: Direct visual confirmation of atom-to-density fit.
- Map Loading:
Tools > X-ray > Get Electron Density Map[00:04:20] - Contouring:
Tools > X-ray > Contour Electron Density Map[00:06:12]- 2Fo-Fc Map: Validates current atom placement [00:04:33]
- Fo-Fc Map: Positive (Green) shows missing atoms; Negative (Red) shows modeled atoms that don't belong [00:04:48]
- Sigma Control: Use '+' and '-' keys. Features should be visible at 1.0–1.5 Sigma [00:05:11]
IV. Handling of Alternative Residue Confirmations
Objective: Understand how to work with alternate residue conformations and save them as separate objects.
- Read in PDB > Search Tab > Enter "1hmt"
- Identification: In high-resolution structures, atoms for a single residue may have two positions labeled 'a' and 'b' [00:08:03]
- ICM Selection Detail: Residues appear with multiple atomic coordinates for the same ID. Toggle or select individually using the 'A' (Atom) selection level [00:08:51]
- The "Clone and Clean" Workflow:
- Right-click the residue >
Clone. Name itState_A[00:09:10] - In
State_A, select the 'B' atoms > Right-click >Delete Atom[00:09:21] - Repeat the process for
State_B, deleting the 'A' atoms.
- Right-click the residue >
- Outcome: This allows for Ensemble Docking, screening ligands against multiple conformations simultaneously [00:09:37]
V. Symmetry & Biological Reconstruction
Objective: Model the functional unit, not just the crystallographic unit.
- Read in PDB > Search Tab > Enter "1cdg"
- Crystallographic Neighbors: Generate the surrounding crystal lattice pocket via
Tools > X-ray > Crystallographic Neighbor[00:10:36] - Read in PDB > Search Tab > Enter "1al2"
- Biomolecule Generator: Reconstruct full assemblies (e.g., viral capsids) via
Tools > X-ray > Biomolecule Generator[00:12:05]
| Prev Introduction to ICM | Home Up | Next Protein Modeling and Engineering |