|Video Example|
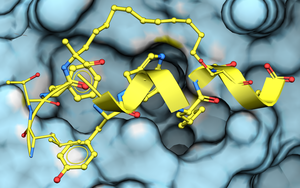
To dock a peptide to a protein structure:
1. Setup the Docking Project and Receptor
- Setup the protein receptor the same way you would for small molecule docking.
- Prepare a table with your peptide sequence(s) to dock from Docking/Dock Peptide table.
2. Preparing a Table with Peptide Sequences
To create a table for peptide sequences, follow these steps:
Create a New Table
- Go to File > New and select the Table tab.
- Choose one string column and set the number of rows based on how many peptides you wish to dock.
- A new table will appear in the GUI.
- Rename the column containing the sequence to 'sequence' .
- Enter your peptide sequences in the format shown below:
Sequence Format Guide
Before defining sequences, check icmff.res in $ICMHOME (distribution directory) to see which residue types ICM supports. All defined residues are stored in the icmff.res file in the ICM distribution directory.
It is a text file so you can view it in a text editor or grep it. Example to search for ornithine grep ornithine icmff.res and you will see the code is 'orn'
1. Standard and Advanced Numbering
- 3-Letter Codes:
gly ala ser pro tyr his - Advanced Notation: (Includes numbering, N/C-terminals, and D-amino acids)
0 nter 1 gly 2 ala 2A Dglu 4 asp cooh
2. Specialized Bonds and Cyclization
- Disulfide Syntax: Use numerical markers in parentheses. Multiple disulfides can be encoded as (1), (2), etc.
acet ala cys(1) ala ala cys(1) ala conh - N-term to C-term Cycles: Use special "virtual" residues:
nvtr ala ala ala ala cvtr
3. Crosslinks and Modifiers
Use curly braces {} for crosslinks or unusual modifications.
- Lys to Glu Link: The 'Xa' stands for crosslink 'a' replacing an atom (e.g., Nz of lysine).
acet ala lys{nz_Xa} ala ala ala ala ala gln{he22_Xa} conh - Unusual Residues:
aba{hg1_C} ser{hg_C(=O)C)}
Note: Modifiers cannot be applied to the backbone (C, Calpha, and N atoms). Hydrogen and side-chain atoms are acceptable.The terminal amide has to be preserved as backbone but you can create terminal amide to side-chain crosslink via a terminal ‘residue’
e.g.
… lys{ce_Xa} …. trp cmet{hm1_Xa}
4. Terminal Designations
| Terminal Type | ICM Residue Code |
|---|---|
| C-terminal amide | conh |
| N-terminal charged amine | nh3+ |
| N-terminal neutral amine | nter |
Example Input File Sequence:
nh3+ nle ala ala ala ala conh
Importing Sequences (Optional)
- You can import a table from an Excel or CSV file. Make sure that the column containing the sequence is labeled 'sequence'.
- If you have a peptide loaded from a PDB file, extract its sequence:
- Right-click on the peptide.
- Select>Extract Sequences to Table.
3. To dock the ligand:
Docking a Peptide Table
Start Docking
- Setup the protein receptor the same way you would for small molecule docking.
- Go to Docking > Dock Peptide Table.
- Click OK to use the default docking settings.
Adjusting Docking Parameters (Optional)
- If you uncheck "dock immediately," you can modify settings before docking.
- Use the table side panel to adjust:
- Thoroughness
- Number of conformations
Restraining or Biasing the Peptide Structure
You can restrain or bias the ligand towards a specific secondary structure. To do this, define each residue with one of the following secondary structure types:
- H - Alpha helix
- G - 3/10 helix
- I - Pi helix
- E - Beta strand
- B - Beta-bridge
- _ - Undefined
- C - Coil