| Available in the following product(s): Homology |
Other resources: Tutorial
Background
Building an accurate model of a loop is very tough. However with small loops ICM has been very successful. ICM was used to design two new 7 residue loops and in both cases the designs were successful. Moreover, the predicted conformations turned out to be exactly right (accuracy of 0.5A RMSD) after the crystallographic structures of the designed proteins were determined in Rik Wierenga's lab (Thanki et al Protein Eng. 1997 10, 159-167). In 2010, scientists at MolSoft developed a a new physics-based internal coordinate mechanices force field (Arnautova et al Proteins. 2010, 79:477) and was tested on loop structures. The new force-field contains new parametization for the dielectric constant, an improved hydrogen bond determination method, and implementation of novel backbone atom torsional potentials which include bond angles of the carbon (alpha) atoms into the internal variable set.There are two loop modeling options:
- Interactive This option will re-build a loop by searching icm.lps contains loop conformations from the PDB database. It searches for suitable loops with matching loop ends and as close loop sequence as possible, inserts them into the model and modifies the side-chains according to the model sequence.
- High Precision Sampling in Batch This option does a full simulation of the loop in the latest ICM force-field as benchmarked in this paper Arnautova et al Proteins. 2010, 79:477. This is a more rigorous simulation than the interactive approach and therefore takes longer to run (see Tutorial here).
Interative - fast loop sampling:
- Read your modeled structure into ICM. Or continue immediately after using build model.
- Select the loop region you wish to model (green crosses in the graphical display). Make sure the object you are selecting is the current object. You can make an object current by right clicking on the object in the ICM Workspace and select "Set to Current"
- MolMechanics/Sample Loop.
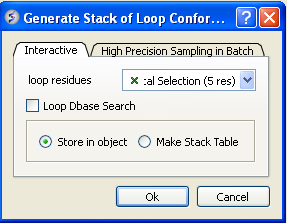
- Select the tab labeled Interactive
- You should see graphical selection in the "loop residue" panel or you can enter the selection here using the icm selection language.
- If you want to perform a fast knowledge-based loop search (searching a database of loops) check Loop Dbase Search.If you would like to perform an ab initio search that takes into account the surrounding environment e.g. a ligand leave the box unchecked. The ab initio search is slower.
- Check whether you would like the stack of solutions saved in an object or in a table. If you choose Make Stack Table a table will be displayed with the loop structures ranked by energy. To view each structure double click on the table. The first row is the loop with the best energy, you can sort a column by right clicking on the column header and choosing sort. If you choose store in an object you can view each member of the stack by clicking on the "+" button in the stack level of the object in the ICM Workspace.
High Precision Sampling in Batch
- Read your modeled structure into ICM. Or continue immediately after using build model.
- Select the loop region you wish to model (green crosses in the graphical display). Make sure the object you are selecting is the current object. You can make an object current by right clicking on the object in the ICM Workspace and select "Set to Current"
- MolMechanics/Sample Loop.
- Select the tab labeled High Precision Sampling in Batch
- Enter a value for effort factor (former 'thoroughness'). This will determine how long the simulation will last.
- Check whether or not you want to sample surrounding sidechains.
- Check whether or not you want to truncate remote parts of the protein.