[ Pocketome | ChEMBL | SureChEMBL | BLAST | UniProt | Ligand Code | Drug Bank | Search PubChem | Historeceptomics ]
| Available in the following product(s): ICM-Browser | ICM-Browser-Pro | ICM-Pro | ICM-Chemist |
The Search tab allows you to search and download content from the following databases:
- The Protein Databank www.wwpdb.org - search for protein and chemical
- The Pocketome Database http://pocketome.org/
- BLAST search the NCBI sequence database
- The UniProt Sequence database http://www.uniprot.org/
- Search the PDB by ligand code.
- Search Drug Bank http://www.drugbank.ca/
- Search PubChem https://pubchem.ncbi.nlm.nih.gov/
- Search ChEMBL https://www.ebi.ac.uk/chembl/
- Search SureChEMBL https://www.surechembl.org/search/
- Search the Crystallography Open Database http://www.crystallography.net/
- Search the HistoReceptomics data from GeneCentrix http://www.historeceptomics.com/
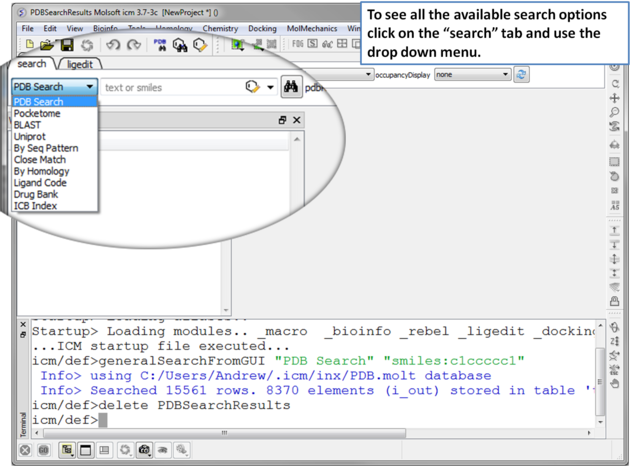 |
| Click on the search tab and then use the drop down menu to identify the database you would like to search and download from. |
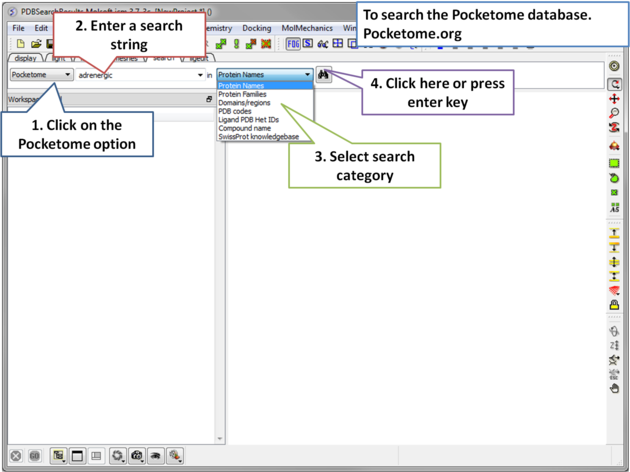 |
| Step 1: Click on the search tab and choose the Pocketome option. Check a field you would like to search e.g. Protein Name, Family, domain... |
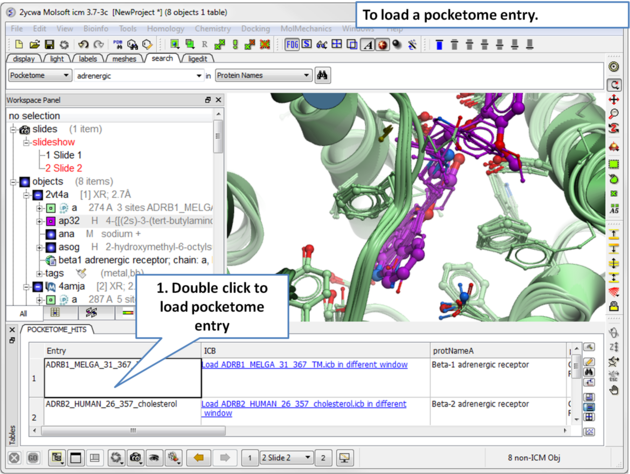 |
| Step 2: A table of pocketome hits will be displayed. Browse the results and double click to load a pocketome entry. |
To search and download data from ChEMBL:
- Click on the Search tab.
- Choose ChEMBL from the drop down button.
- Enter a search string or search by chemical sketch by clicking on the editor button in the panel.
- Select a field to search e.g. Protein Name.
- Select whether you wish to return just activity data or all data.
- Click the search button.
A table will be displayed as shown below.
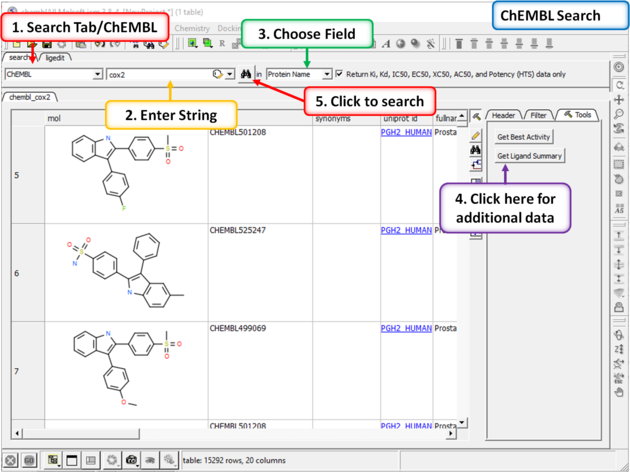
The table contains the original activity data without merging or aggregation.
For instance,
Chemical 1, Protein 1, Activity 1
Chemical 1, Protein 1, Activity 2
Chemical 1, Protein 2, Activity 1
Chemical 2, Protein 1, Activity 1
Click on the button Get Best Activity to display the best activity for each chemical, protein pair.
For instance,
Chemical 1, Protein 1, Best(Activity 1, 2, 3, ...)
Chemical 1, Protein 2, Best(Activity 1, 2, 3, ...)
Chemical 2, Protein 1, Best(Activity 1, 2, 3, ...)
Click on the button Get Ligand Summary to display a list of chemicals with all activities found in ChEMBL.
For instance,
Chemical 1, Protein 1, Activity 1
Protein 1, Activity 2
Protein 2, Activity 1
Protein 3, Activity 1
Chemical 2 Protein 1, Activity 1
Protein 2, Activity 1
Protein 2, Activity 2
- Click on the Search tab.
- Choose SureChEMBL from the drop down button.
- Enter a search string or search by chemical sketch by clicking on the editor button in the panel.
- Select a field to search e.g. Protein Name.
- Select whether you wish to return just activity data or all data.
- Click the search button.
To BLAST search the NCBI sequence database:
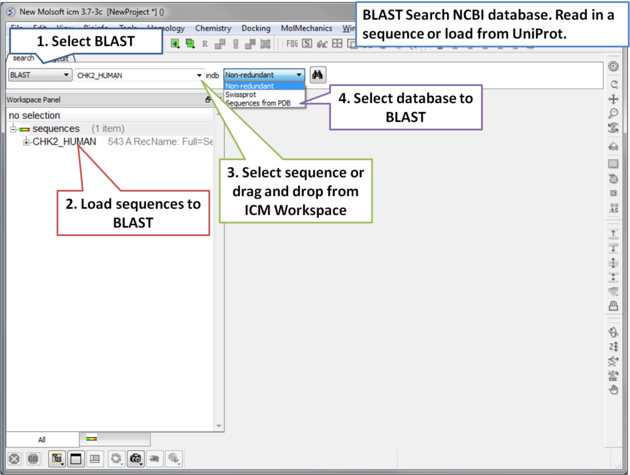 |
| Step 1: Load a sequence into ICM. Select the Search tab and choose the BLAST option. Drag and drop the sequence into the search field, use the drop down menu or type the sequence name. |
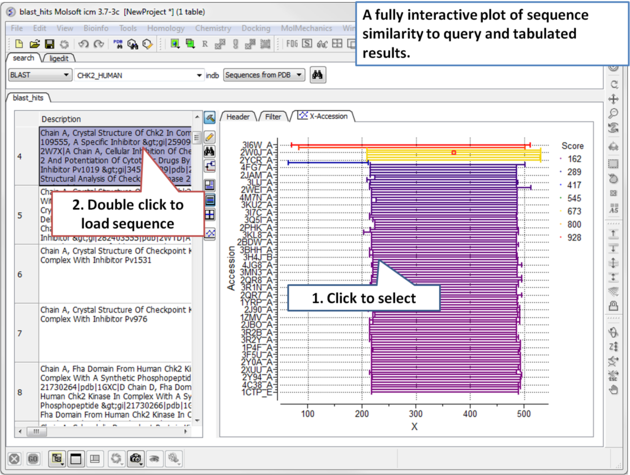 |
| Step 2: A fully interactive table and plot of sequence conservation will be displayed. Double click to load a sequence. |
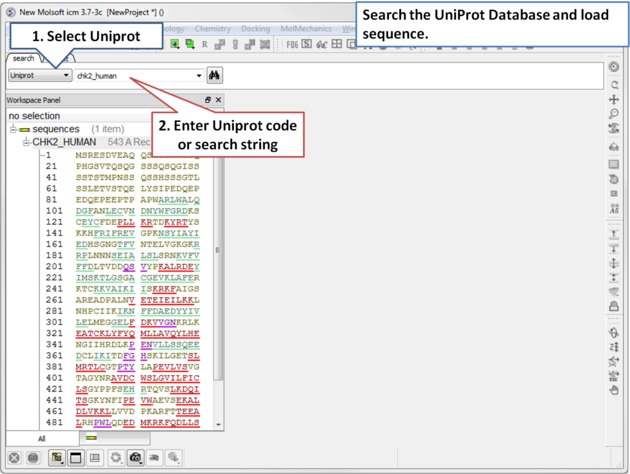 |
| Step 1: Click on the search tab and select Uniprot from the drop down menu. Enter the UniProt code. The sequence will be loaded directly into ICM and you will see it in the ICM workspace. |
To find structures containing a particular ligand in the PDB.
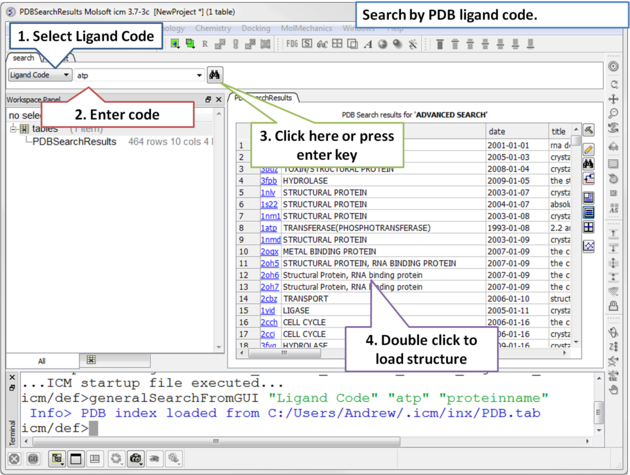 |
| Step 1: Click on the search tab and select Ligand Code from the drop down menu. Enter the Ligand code and a table of hits will be displayed. |
To search Drug Bank ( http://www.drugbank.ca/)
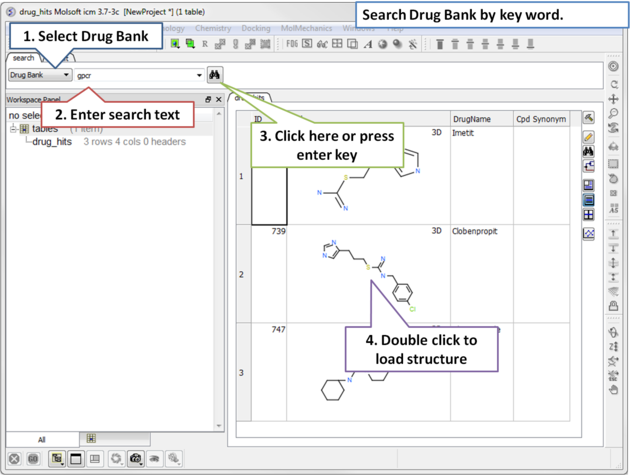 |
| Step 1: Click on the search tab and select Drug Bank from the drop down menu. Enter a search string and a table of results will be displayed. |
The table contains the original activity data without merging or aggregation.
For instance,
Chemical 1, Protein 1, Activity 1
Chemical 1, Protein 1, Activity 2
Chemical 1, Protein 2, Activity 1
Chemical 2, Protein 1, Activity 1
Click on the button Get Ligand Summary to display a list of chemicals with all activities found in Drug Bank. The button is located in the extra panel (click on hammer - top right of table panel).
For instance,
Chemical 1, Protein 1, Activity 1
Protein 1, Activity 2
Protein 2, Activity 1
Protein 3, Activity 1
Chemical 2 Protein 1, Activity 1
Protein 2, Activity 1
Protein 2, Activity 2
To search PubChem (https://pubchem.ncbi.nlm.nih.gov/):
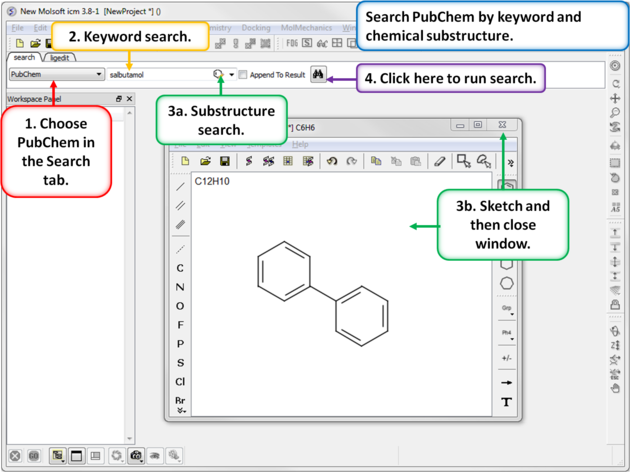 |
|
The historeceptomics (HR) platform is developed by GeneCentrix and reports tissue specific profiling for a drug molecule. It is a unique bioinformatics platform designed to identify mechanism of activity and predict unforeseen effects a chemical might have in the target tissue and off-target tissues.
HR integrates data from a variety of sources including:
- BioGPS
- GTex
- ChEMBL
- DrugBank
- BindingDB
- Genecentrix database
- Select the Search tab.
- Select the historeceptormics option from the drop down search menu.
- Enter a drug name or sketch a chemical using the molecular editor.
- Choose the BioGPS or GTex database. Selecting BioGPS will use the GeneAtlas U133A dataset from BioGPS to analyze the tissue-wide expression profiles of targets of interest. Selecting GTEx will use the median gene-level TPM by tissue dataset from GTEx to analyze the tissue-wide expression profiles of targets of interest
- Click the search button.

After searching a results panel as shown below will be displayed.
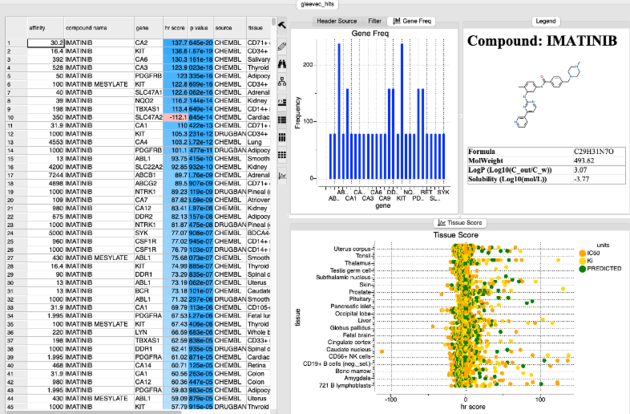
The table is ranked by hr Score and reports the binding affinity gene name and tissue. The p-value represents the chance that the overabundance of the tissue/target in question could occur by random chance based on the variability of the expression of that target across 70 other tissues. Additional plots of gene frequency and hr score for each tissue are reported.